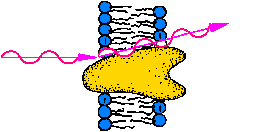
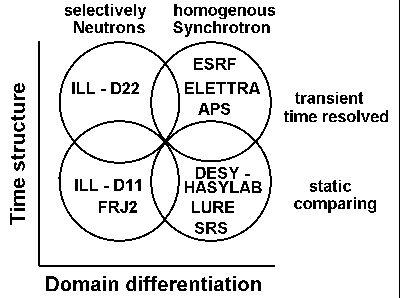
| Inst. f. Biochemie, Universität Mainz, AG Membranenstruktur -
Protein Dynamik |
PD. Dr. T. Nawroth |
| Max-Planck-Institut für Biochemie, Martinsried, Biophys. |
PD. Dr. H. Heumann |
| Inst. für molekulare Biophysik, Universität Mainz |
Prof. Dr. H. Decker |
| Inst. f. Biochemie, Universität Mainz, MolekularBiologie |
Prof. Dr. C. Koch-Brandt, PD. Dr. G. Klock |
| Physics Department, Institute E13, Technische Universität München
TUM, Garching |
PD. Dr. Wolfgang Doster, Prof. Dr. W. Petry |
| Physics Department, Institute E17, Technische Universität München
TUM, Garching |
Prof. Dr. F. Parak |
| Max-Planck-Arbeitsgruppen Strukturelle Biologie, c./o. DESY-HASYLAB,
Hamburg |
Prof. Dr. E. Mandelkow |
| GKSS Forschungszentrum, Neutronenstreuung, Geesthacht |
Dr. R. Willumeit |
.