 |
Membrane Pharmacy Structure Dynamics(MPSD)
Research group : Priv.Doz. Dr.
Thomas Nawroth
|
|
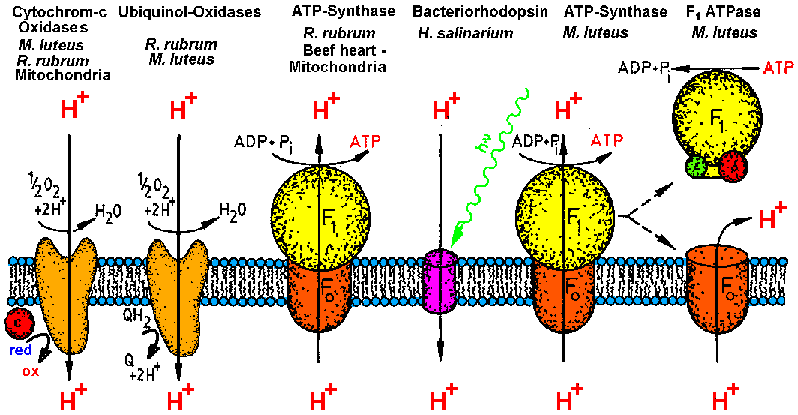
| Membrane proteins investigated by the MPSD group in the last ten years.
Currently the work is focussed on bacterial ATP-synthase, its catalytic
head fragment F1ATPase and Cytochrome-c oxidase COX (from [T20]). |
Membranes are hetrogenous complexes of lipid bilayers, membrane proteins
and other macromolecules (carbohydrates, cell sceleton elements). The biological
function depends strongly on structure and
the moleclar action between lipids and proteins. The investigation of structure
and function requires a concept of isolation,
purification and reconstitution yielding model membrane systems. This can
be most easily done with bacterial systems, which can be cultured in large
amounts and manipulated, e.g. by moleculare genetics. Because of the complex
dependence between structure and function the investigation of membrane
proteins requires the parallel application of methods of structural biology,
biophysics and biochemistry (functional analysis). This is the interdisciplinary
concept
of biophysical chemistry.
The energy metabolism of most cells (bioenergetics)
depends on ion pumping membrane proteins, which are proton pumps in most
recent organisms but sodium or other ion translocating proteins in archeae
(archaebacteria) and extremophiles. The reason for this is the intrinsic
flexibilty of membrane protein structure, which is a consequence of the
hydrophobic effect. This results in a binding energy by stripping of hydration
water during membrane insertion of hydrophobic molecules but not in a directed
force between the membrane components. Thus membrane components can move
in the membrane plane with minimal energy input (lateral translation or
rotation).
The isolation and purification of the
membrane proteins requires the solubilization of the protein by replacement
of the lipids by detergents (both are amphiphiles). As shown in the below
figure, the native structure and thus the biological function is only retained
if weak, non-denaturating detergents of highest chemical quality are used
(no peroxides or long chain impurities). The methods and technology of
membrane protein purification is subject of the lecture
"Methods of membrane biochemistry". The methods for structure research
of membranes and membrane proteins in solution are subjects of the lecture
"Biophysical chemistry of membranes and membrane proteins". You can learn
the technology of membrane protein purification and some techniques for
biophysical chararacterization by participation in our practical
course "Advanced practicum in membrane biochemistry".
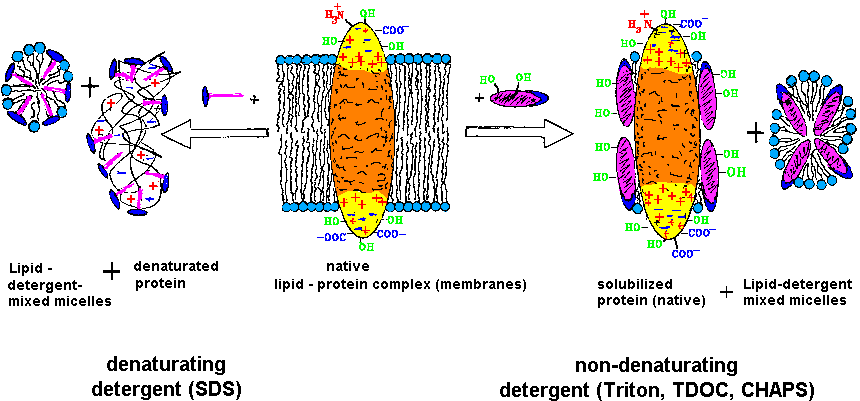 |
| The purification and investigation of membrane proteins requires the
retainment of the function competent structure (native), which is obtained
by solubilzation with weak, non-denaturating detergents [T20]. |
.
The membrane structure dynamics group (MPSD) has during the last decade
investigated the proteins shown in the first picture. The scientific
concept is described explicitly in [T20].
The results of 50 man-years research are described in the thesis
list, several publications, applications
and reports. The objects are the following
proton pumps, which are capable of molecular motions:
ATP-synthase : The enzyme (M=500,000)
converts the energy of the ion transmembrane transport to that stored in
the energy rich compound ATP during molecular rearrangements. F1ATPase
is the catalytic head fragment of ATP-synthase (M=400.000). The pumped
ions are mostly protons, but as found by Prof. P. Dimroth, in some cases
sodium ions are transported. The static structure of an ADP-inhibited modification
of F1ATPase was resolved with atomar resolution by X-ray crystallography
(Nobel avard: J.E: Walker 1997). Nevertheless the reaction mechanism is
essentially unknown. The reson is the lack of time resolution in the very
most studies. We investigate molecular motion inside ATP-synthase and F1ATPase
by time resolved X-ray small angle scattering after activation by rapid
mixing methods (stopped-flow) or flash photochemistry
(caged ATP, caged acids - proton). This yields a structural
film of working ATP-synthase. The structural dynamics upon molecular
regulation of ATP-synthase are investigated by comparative static X-ray
small angle scattering. The structure of reconstituted
ATP-synthase in situ is studied by Neutron small angle scattering of
contrast matched liposomes.
Cytochrome-oxidase COX, Quinol-oxidaseQOX
(Cytochrome-o) and Cytochrome-c
reductase (Cytochrome-bc1 complex) are
Cytochrome-group containing oxido-reductases, which combine an oxidation
of external coenzymes (ubiquinol, cytochrome-c) with the proton membrane
transport in oxidative phosphorylation or photosynthesis (bacteria). We
investigate molecular motions of Cytochrome-c oxidase COX and Quinol-oxidase
QOX [F19] from Micrococcus luteus
during regulation by comparative X-ray small angle scattering of enzyme
modifications (oxidized, reduced, inhibited ...). Structural dynamics have
been shown in cooperation with PD. Dr. W. Doster
by time resolved optical spectroscopy after flash irradiation of carbon
monoxide (CO) inhibited Cytochrome-c oxidase COX from Micrococcus luteus.
In cooperation with PD. Dr. T. A. Link this
was compared to Cytochrome-c oxidase from beef heart mitochondria. The
Cytochrom-o complex [F12] and the Cytochrome-bc1
complex (Cytochrome-c reductase) from Rhodospirillum rubrum have
been investigated by biochemical methods.
Bacteriorhodopsin is a light
driven proton pump and the parent of the 7-helix receptor protein family.
The protein of 27,000 mass burries a retinal chromophor in the center of
a hollow protein cylinder, which consists of seven amphipatc helices (the
outer side is hydrophobic). In Halobactium salinarium (formerly
called Halobacterium halobium) the protein is arranged in natural
2D-crystals, which consist of a hexagonal arrangement of Bacteriorhodopsin
trimers and 15% membrane spanning lipids. These specialized membrane domains
are called "purple membrane" PM, which is
the operative unit of halobacterial photosynthesis. Bacteriorhodopsin is
extremely stable: during our ERA sub-experiment [F21],
purple membrane samples survived a 6 month flight in space (extreme vacuum
an cold, dark (no sun light) samples). For biochemical and structural investigations
we solubilize the purple membrane by detergent and purify the otained monomeric
Bacteriorhodopsin mBR bei HPLC [F5].
The stzucture of mBR in detergent solution is invetigated by neutron small
angle scattering, X-ray small angle scattering and time resolved flash-photolabeling
using our 3-domain photolabels (photoreactive, selective, detectable).
Those "lantern-labels" are available with metal-heads, detectable with
anomalous X-ray scattering (Eu, Tb), Mössbauer spectroscopy (Fe) or
NMR (Gd), and with fluorescent or colored head groups (dansyl, dabsyl ...)
[F15]. The labels can be introduced in
polymers for the detection of molecular motions in "motile polymers".
.
Reconstitution
of membrane proteins in model membranes
|
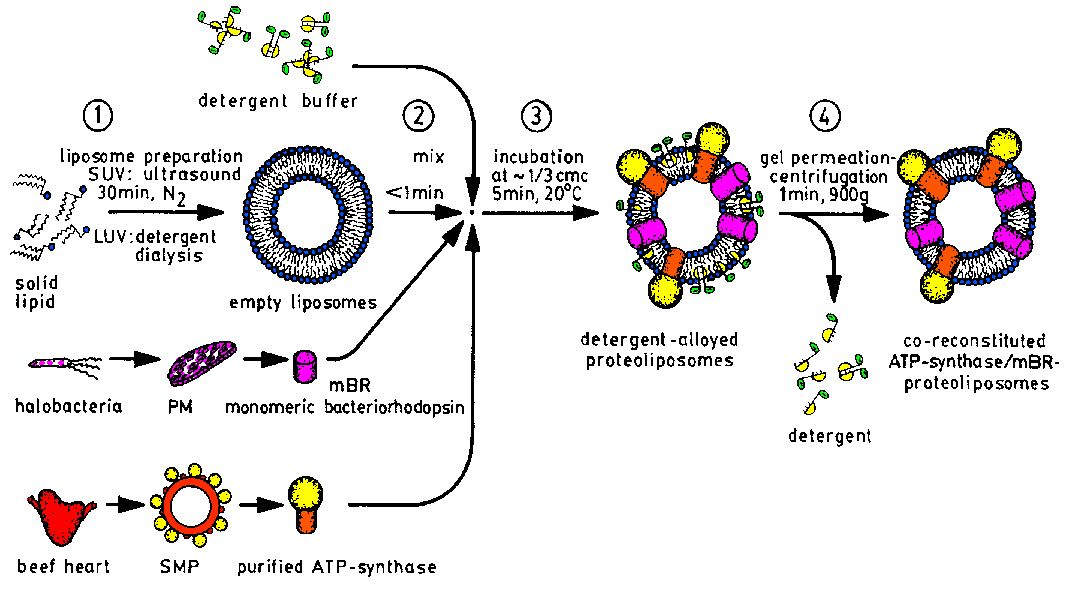
| The reconstitution of purified membrane proteins into liposomes as
model membranes is done by our "detergent incubation of preformed liposomes"
procedure [F3]. Proteoliposomes for functional
analysis [F16] may contain several membrane
proteins [F17,F18,F22,F23],
whereas those for structue investigation contain only one protein molecule
in small unilamellar vesicles (SUV) [F3,F9]. |
.
The investigation of the purified membrane proteins is done at two
levels: i) investigation of structure and function of the detergent solubilized
mebrane protein, and ii) investigation after reconstitution into model
membranes. As model membranes we use liposomes and BLM (black lipid membranes,
planar membranes of 1mm diameter). For the structural study of protein-
detergent solutions we use only those detergents, which do not disturb
the scattering signal of proteins by micelle scattering [F6].
These are special non-denaturating detergents, which form only very small
micelles, e.g. TDOC, CHAPS,Octylglucoside. The reconstitution of purified
membrane proteins into liposomes as model membranes is done by our "detergent
incubation of preformed liposomes" procedure [F3].
During reconstitution detergent alloyed liposomes are formed, which are
in case of charged detergents stabilized by electrostatic repulsion according
to neutron small angle scattering experiments [F7].
For functional analysis [F16] the obtained
proteo-liposomes may contain two coupled membrane proteins in several copies
in large liposomes, e.g. monomeric bacteriorhodopsin and ATP-synthase [F17,F18,F22,F23].
Proteoliposomes for structue investigation by neutron small angle scattering
of contrast mached liposomes must contain only one protein molecule in
small unilamellar vesicles (SUV) [F3,F9]
to avoid interference signals between lipid density fluctuation and protein
scattering amplitudes [F3,T1].
The liposomes can also consist of monomeric or polymeric polymerizable
lipids containing butadiene or di-ecetylene groups, as investigated in
cooperation with Prof. H. Ringsdorf [F11].
The surface exposition and distances in and between subunits of membrane
proteins are investigated by flash photolabeling.
email to: nawroth@MPSD.de
update : 15.10.2013




